About
The project
The fly cancer screen is a Cancer Research UK supported project, which uses the fruit fly, Drosophila melanogaster, to identify conserved genes that enhance or suppress epithelial tumour progression.
Publication
Further information can be found in the following publication:
A genetic analysis of tumour progression in Drosophila identifies the cohesin complex as a suppressor of individual and collective cell invasion
Brenda Canales Coutiño, Zoe E. Cornhill, Africa Couto, Natalie A. Mack, Alexandra D. Rusu, Usha Nagarajan, Yuen Ngan Fan, Marina R. Hadjicharalambous, Marcos Castellanos Uribe, Amy Burrows, Anbarasu Lourdusamy, Ruman Rahman, Sean T. May, Marios Georgiou
ISCIENCE (2020): https://www.cell.com/iscience/fulltext/S2589-0042(20)30422-3
Metastasis is the leading cause of death for cancer patients. Consequently it is imperative that we improve our understanding of the molecular mechanisms that underlie progression of tumour growth towards malignancy.
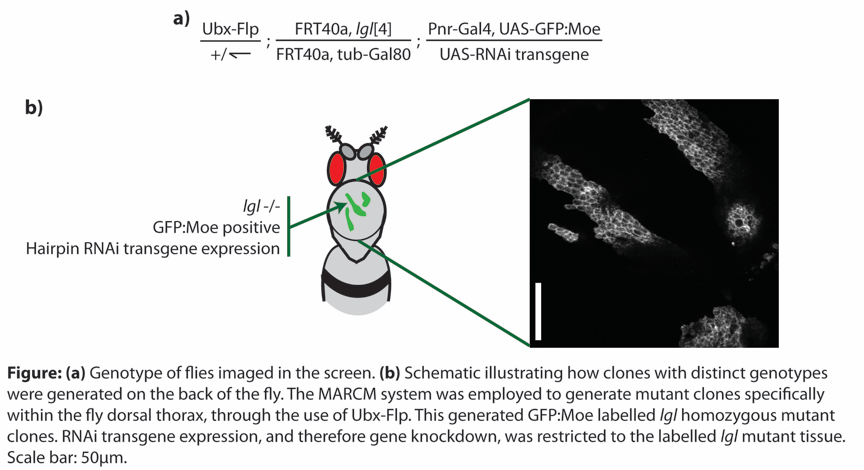
We have developed a system by which we can study epithelial cell shape in high temporal and spatial resolution in the living animal. This in vivo genetic system allows us to: (1) generate a patch of tissue on the dorsal thorax that is homozygous mutant for a tumour suppressor, surrounded by wild-type tissue; (2) specifically label the mutant tissue with GFP:Moe (the actin binding domain of moesin fused to GFP), thereby labelling the actin cytoskeleton of these cells; (3) overexpress an RNAi transgene to deplete expression of a gene of interest specifically within the mutant, labelled tissue (see Figure). By combining the genetic amenability of Drosophila melanogaster with the power of RNAi transgenics, we were able to generate tumours with specific genotypes and to monitor tumour behaviour in the living animal.
Using this system we screened through 764 RNAi lines, corresponding to 497 Drosophila genes, whose human orthologues have been found to be commonly mutated or misregulated in a variety of human cancers. This screen has made a number of novel findings:
- We identify numerous novel conserved genes that either positively or negatively regulate specific aspects of tumour behaviour within an epithelium in a living animal.
- We identify a number of conserved ‘invasion suppressors’ that promote tumour cell invasion upon loss of expression. We additionally generate a functionally validated network of invasive genes, which includes 140 genes, 99 of which have not been previously implicated in cancer cell invasion or migration.
- Our detailed analysis of how conserved candidate genes affect tumour behaviour in a living animal has generated a data set that provides a first step in the characterisation of hundreds of genes that are commonly mutated or misregulated in human cancers.
- This work establishes the fly pupal notum as an excellent in vivo system to study tumour progression.
We have generated a database with 33 different phenotypic categories, each describing an aspect of tumour behaviour (e.g. invasion, multilayering, tumour size, proliferation, apoptosis, junction defects, abnormal cell shape, etc.). This systematic scoring of specific aspects of tumour behaviour allowed us to record a detailed analysis of how each gene knockdown affected tumour behaviour.
To understand tumour transition to malignancy, and to develop new therapeutic strategies, it will be key to understand the molecular basis of this event. This work enabled the discovery of functional cooperative regulators of invasion and the establishment of a network of conserved invasion suppressors, which will help to inform future research in this area.
Acknowledgements and who we are
This work was supported by a Cancer Research UK Career Establishment Award [grant numbers C36430, A12891], with additional funding from CONACYT and a Nottingham Vice-Chancellor’s Scholarship for Research Excellence Award.
The project was carried out over a number of years by a team of researchers at the Georgiou lab, University of Nottingham, School of Life Sciences. Researchers that made major contributions to the design and completion of the screen include: Brenda Canales Coutiño, Zoe E. Cornhill, Africa Couto, Natalie A. Mack, Alexandra D. Rusu, Usha Nagarajan, Yuen Ngan Fan, Marina R. Hadjicharalambous, Amy Burrows and Marios Georgiou, with additional technical support from Zsuzsa Markus.
The Georgiou lab uses the fruit fly, Drosophila melanogaster, to study cell morphology and cancer biology. Research at the lab lies at an interface between cell, developmental and cancer biology, with recent work addressing a number of fundamental processes such as the regulation of the actin cytoskeleton, cell polarity, cell morphology, cell adhesion, endocytosis, trafficking, tissue architecture and cell signalling.
You can contact the team by emailing marios.georgiou@nottingham.ac.uk
We would like to thank the fly community for their generosity with reagents, the Bloomington, VDRC and NIG stock centres, without whom this project would not have been possible, and SLIM (School of Life Sciences Imaging) for invaluable help with the confocal microscopes. We additionally thank Keith Spriggs for help with preparing image files for the online resource.
